
Initial Breed Summary: Irish Red and White Setter
The Irish Red and White Setter is a small breed of setter that was revived in the 20th century post World War I by Reverend Noble Huston of Ballynahinch and other breeders; later in 1940, Mrs. Will Cuddy nursed a female puppy to health. This female was named “Judith Cunningham of Knockalla” and is behind every Irish Red and White Setter alive today . What has UC Davis and esteemed Dr. Niels Pedersen DVM PhD found in regards to the breed? You can find the full report here.
Inbreeding – how genetically similar are the parents of typical Irish Red and White Setter?
Dr. Pedersen found the following in terms of the inbreeding values of this population:
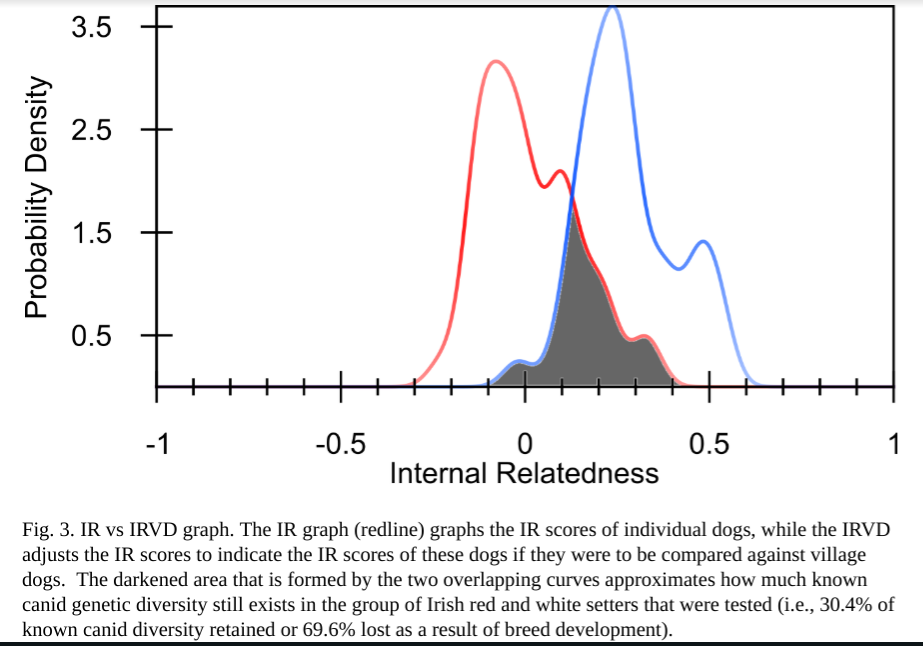
The entire population is on average slightly inbred with a mean IR score of 0.014 (Table 4, Fig. 3). However, one half of the dogs had IR values from -0.081 to +0.207 and one fourth from 0.103 to 0.338. An IR score of 0.103 would be equivalent to mating half siblings from a random bred population, while an IR score of 0.388 would be equivalent to breeding full siblings from a highly inbred population. This inbred population was countered with an equal number of dogs with IR scores as low as -0.084 to -0.224.
Interpretation? A significant portion of this population is highly inbred and could benefit from predictive software to aid in choosing unrelated (as possible) breeding mates.
However, being homozygous or heterozygous, or inbred or outbred, is not an indication of biodiversity of the breed.
Breedwide diversity
Biodiversity and allelic richness are terms we try to discuss regularly at BetterBred. Namely, how many different versions of genes or alleles exist in each breed? Breeds with more allelic richness and whose genetics are also well distributed throughout the population tend to be healthier. So what di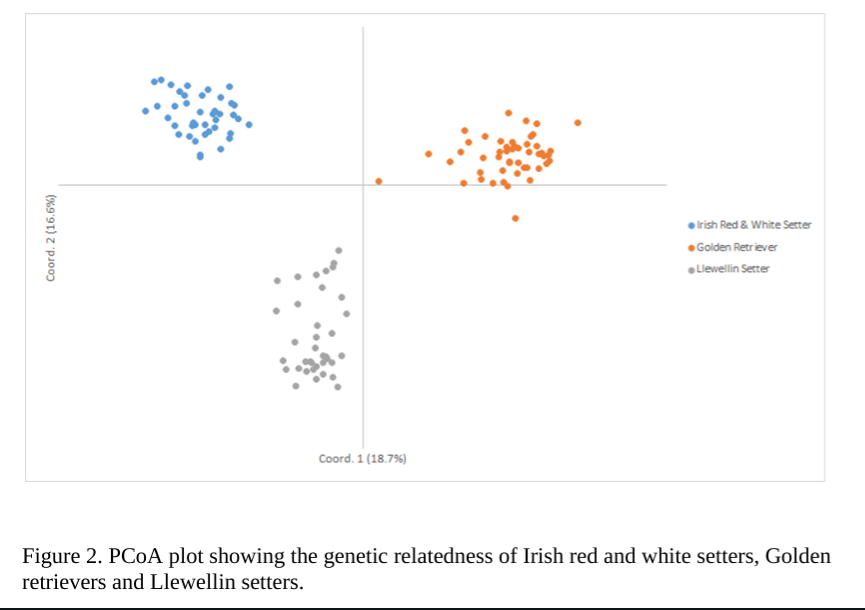 d they find about the Irish Red and White Setter population?
d they find about the Irish Red and White Setter population?
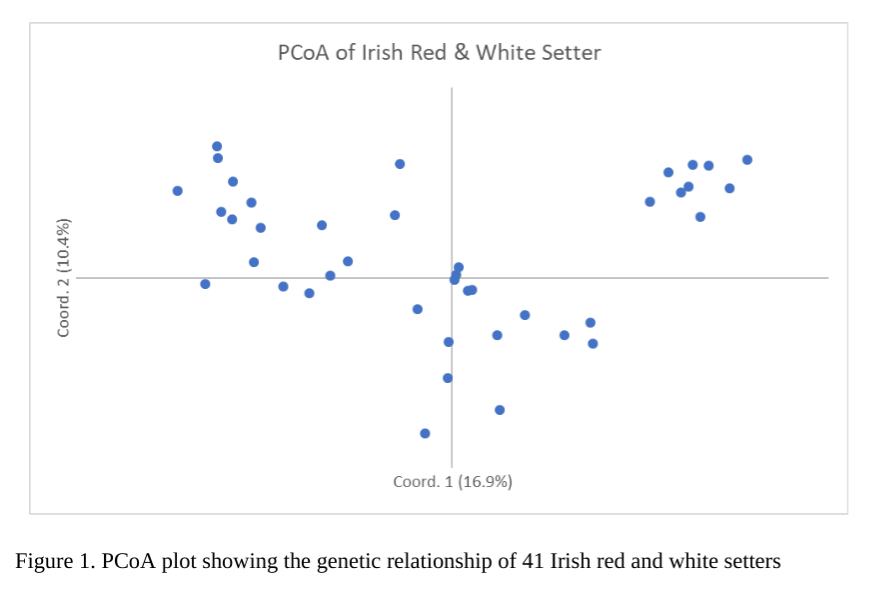
One of the first things you can notice is that Irish Red and White Setter form a single breed with some clustering within the breed, as seen by the PCoA graph above (figure 1 above).
Dr. Pedersen writes, “(the) PCoA plot of the 41 Irish Red and White Setters that shows them clustering as a single breed, as would be expected. However, individuals are not tightly clustered around the central X/Y axis but are spread at some distance from each other across the graph. This suggests that this group of dogs is genetically diverse, and this diversity may reflect phenotypic differences as well. A group of nine dogs is clustered as outliers in the upper right quadrant and probably represent closely related dogs from the same kennel or bloodline. ”
When the PCoA is made to include several breeds, you can see that the breed is distinct from the Llewelin Setter and Golden Retriever. It will be very interesting to see how they may cluster with the Irish Red Setter, should they be tested in the future.
But what about biodiversity or allelic richness? Irish Red and White Setters have low diversity in comparison to other tested breeds, but not the lowest of breeds tested to date. Breeds with lower biodiversity include Alaskan Klee Kai, Swedish Vallhund and Shiloh Shepherds. However, as more are tested it is likely more biodiversity will be found.
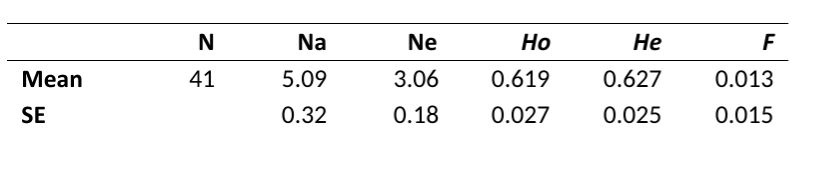
Average alleles per locus and effective alleles per locus are aspects that we discuss in these breed reports. The closer the effective alleles (we call these “alleles that are effectively contributing to your population”) are to the average alleles per locus, the better distributed the genetics in the breed are. Why would you want well distributed genetics? It keeps risk of unwanted recessive traits from appearing, lowers risk of random loss of diversity (called genetic drift) and it lowers the frequency of both simple and complex disease haplotypes that might unintentionally become more common if there were a bottleneck. The average alleles per locus in this breed was found to be 5.09, while the effective alleles per locus was found to be 3.06.
What does this mean? The effective alleles are closer to the average alleles per locus than is typical for many breeds, meaning – in this tested sample – the Irish Red and White Setter is doing a good job maintaining what they have.
Each breed should make an effort to raise their effective alleles per locus. How do you do this? You use BetterBred’s breed management software to breed for higher than breed average Outlier Index (OI). This measurement was created to help breed communities maintain healthy distribution of breed genetics or redistribute unbalanced diversity, even while they select for the traits they want in their dogs.
Diversity compared to village dogs
Dr. Pedersen’s hallmark comparison is to a combined population of village dogs. These domestic dogs have not been shaped by human effort, but instead have evolved through random breeding and natural selection. They therefore have a great amount of genetic diversity which is representative of the original diversity dogs had prior to modern breeding methods which changed them. Dr Pedersen assesses the IR results based on only the breed’s genetic data and then compares these to their results using the village dog genetics – called IRVD. The difference between these two results illustrates the diversity that has been lost in each modern breed.
From the report:
The IRVD curve for Irish red and white setters was shifted well to the right, reflecting a 69.2% loss (or 30.8% retention) of potential genetic diversity during breed development (Fig. 3).
Translation? Irish Red and White Setters have lost a significant amount of biodiversity in comparison to the other tested breeds to date. However, this may change as more of the breed is sampled worldwide.
DLA – Dog Leukocyte Antigen
A Look at the Dog’s Immune System!
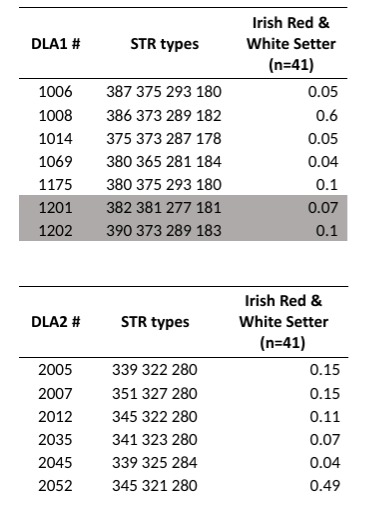 In the 41 Irish Red and White Setters that have been tested to date, there are 7 DLA class I and 6 DLA class II haplotypes discovered. Two Class 1 DLA haplotypes have not yet been identified previously by UC Davis (highlighted in the image from UC Davis to the right).
In the 41 Irish Red and White Setters that have been tested to date, there are 7 DLA class I and 6 DLA class II haplotypes discovered. Two Class 1 DLA haplotypes have not yet been identified previously by UC Davis (highlighted in the image from UC Davis to the right).
According to the report:
The incidence of most of the haplotypes was greater than 4-5%. However, one DLA class I (1008) and one DLA class II (2052) are found in 50-60% of individuals. These haplotypes are likely to remain dominant even after more dogs are tested, although several additional low incidence haplotypes are likely to be identified as more dogs are tested. It appears that contemporary Irish Red and White Setters have evolved from only seven DLA class I and II haplotype lineages. These lineages were most likely descended from dogs accepted by the Irish Kennel Club at the time of the revival of the breed in the 1970s.
It might be interesting to consider that the most common DLA haplotype combination may have descended from “Judith Cunningham of Knockalla” , the female that is behind all IRWS today. In order to preserve the DLA haplotypes existing in this population, breeders should take note of the atypical DLA haplotypes in the population so as not to lose them to genetic drift, or random extinction. You can read more about using DLA in a breeding program here.
The breed report also noted the following:
DLA class I and II haplotype sharing were greatest with the Golden retriever, and secondly with the Standard and Toy Poodle and Giant Schnauzer. Unfortunately, the VGL has no DLA class I and II haplotype data for the Irish Setter with which to compare.
DLA comparisons are always extremely interesting as it can inform on origins of breeds; breeds that have more DLA association indicate evolutionary development of breeds, as opposed to those with fewer DLA haplotypes in common. It will be interesting to see how the Irish Red Setter would compare to the Irish Red and White Setter.
Conclusion
From Dr. Pedersen’s report:
It is not possible based on 41 individual dogs to make definitive conclusions, but the results of initial testing do provide important insights. This initial population of dogs appeared, on average, to be products of parents that were as unrelated as possible. However, IR scores that more accurately measure the relatedness of parents of individual dogs identified a significant sub-population of highly inbred dogs. This might be expected in a small breed with relatively low numbers. It can be difficult to identify least-related mates that are accessible when needed.
The breed appears to have average genetic diversity compared to other breeds, having retained an estimated 35% of the autosomal STR alleles known to currently exist among all dogs. However, the number of founders or closely related founder lines present in contemporary Irish Red and White Setters is small based on the percentage of known canid DLA class I and II haplotypes (i.e., 3-5%). The lack of genetic diversity in the breed has been confirmed from pedigrees, with an effective population size of 39.4 individuals (10).
More biodiversity may be discovered as more dogs are tested. More samples are needed from all around the world.
Dr. Pedersen’s report also notes that the IRWS have a sanctioned outcross program and has the following commentary:
The benefit of using Irish Setter is that much less backcrossing would be required to re-establish the phenotype. On the downside, it may not add much to current genetic diversity. However, the Irish Setter does not have the severe historical genetic bottlenecks suffered by the Irish Red and White Setter and it is possible that they possess much more of the original genetic diversity of the original Irish Setter founders than existing Irish Red and White Setters. The truth can only be determined by intensive DNA testing of both breeds to determine how much new diversity would be added. DNA testing would also determine which dogs from both breeds would be best to use for outcrossing. The introduction of new genotypes will also have to be closely monitored to assure that it is commingled as efficiently as possible with existing genotypes and so that the introduction of new genetic diversity is not done at the expense of existing genetic diversity.
Testing dogs from these outcross programs will be valuable to determine whether they are adding genetic diversity to the breed or if they are genetically similar to the IRWS; tracking descendants and maintaining the infusion of new genetics would also be of value, as new genetic diversity can be rapidly lost with backcrossing to the mainstream population. Maintaining this can be done by using our breed management software to detect which dogs are genetically atypical from the main population (while selecting for breed type).
Our Recommendation
Since the Irish Red and White Setter is a relatively healthy breed, we recommend breeders test their dogs and use the testing to select the most unrelated quality breeding mates possible within this population, as well as maintain the breed’s biodiversity. Outcrossed dogs should be tested to determine if they are adding biodiversity to the breed, and then descendants should be tracked to insure the influx of new genetics is not lost; our breed management software will be of great use for this purpose. Breed for IR at or below zero and OI at or above breed average to breed for outbred puppies and to maintain the breed’s biodiversity. When all else is equal, take note of atypical DLA haplotypes and make an effort to redistribute those that are poorly represented.
More samples are needed from this breed to establish a full breed report. If you have a Irish Red and White Setter and would like to contribute to the research phase, you can do so here.
 Previous Post
Previous Post Next Post
Next Post


