It’s rare that we see Embark attack BetterBred directly by name. They usually (inexplicably) attack only the highly qualified scientific work done by the experienced researchers at UC Davis’ Veterinary Genetics Lab. Considering how often Embark insists their testing methods are superior, it is surprising how often they attempt to undermine confidence in the VGL test or in our little company. I have often been dismayed at their tactics, but I have understood them to be due to the competitive pressures involved with a certain group of entrepreneurs who all want to be the next Mark Zuckerberg, and found the next Facebook or Blue Apron. That pressure breeds a certain distasteful cut-throatedness that has done away with the old fashioned concepts of fair play and collegiality. Unfortunately, this is the world we are in now, and since employees of Embark are finally attacking our methods publicly by name, we are forced to address their claims head on.
I was recently forwarded something from several sources, and it was posted publicly on Facebook as follows:
“From Erin Chu (embarkvet) on:
“Sep 7, 00:19 ADT
“Both BetterBred and Embark include genetic diversity tools to help breeders make the most informed breeding choices. How we actually assess diversity, however, differs greatly. Really the most important thing is the fact that BetterBred bases its analyses off a panel of 33 STRs (simple tandem repeats). There is nothing wrong with using STRs—in fact they are still the mutation class of choice in simple paternity cases and some crime scene investigations. It’s just that 33 of anything would be considered a limited marker panel, which could be perfectly sufficient to resolve a paternity or DNA fingerprinting case, but would largely be considered insufficient for assessing genome-wide diversity in a species with a 3 billion basepair genome. In fact, since dogs have 39 chromosomes, using a 33 STR panel means we don’t even have one STR per chromosome—meaning we’re just skipping over these chromosomes in our diversity assessment. In comparison, Embark measures genetic diversity using over 200,000 SNPs (single nucleotide polymorphisms). This means that we’re looking at genetic diversity at a more than 6000X higher resolution than a 33 STR panel. This allows us to identify very short runs of homozygosity (ROH), which have been shown to harbor deleterious alleles (as my colleague Aaron Sams published on recently in Sams et al, 2018). You miss these ROH altogether with a less dense panel. Is that a bad thing? Well, yes, if your goal is to maintain genome-wide diversity and reduce inbreeding. Further work from Aaron’s group will be published soon and demonstrates that actually, if you select for genetic diversity using just 33 markers, you maintain diversity at those 33 markers, but you LOSE genome-wide diversity at a faster rate than if you just selected mates randomly. Some of his preliminary work on this topic can be read here: https://embarkvet.com/breeding-for-the-future-why-genome-wide-diversity-matters”
Erin Chu is an employee of Embark and therefore not unbiased in her opinions – and she has never had, as far as we are aware, any access to the tools used by BetterBred. She therefore cannot legitimately comment on them, demonstrates no comprehension of how they work, and yet openly and freely mischaracterizes them in her statement. This is both unprofessional and unscientific. I can only imagine that it is Embark’s commercial need to drive out any other innovator from the market that would inspire this unfortunate statement.
The only fact Chu knows to use in her argument against BetterBred is that the Embark panel, which is a repackaged Illumina CanineHD SNP chip, has more individual markers than the UC Davis Veterinary Genetics Lab’s canine diversity test. This is quite an obvious and simplistic observation, and the message seems to work for a while on breeders and pet owners who aren’t geneticists. Embark never explains the qualitative difference (what they are for) between SNPs and STRs, only the quantitative difference (how many there are.) These markers, of course, cannot be compared one to one and are used for very different purposes, which Chu, a veterinary geneticist, ought naturally to know. Fortunately, the more experienced veterinary and conservation geneticists at VGL who developed the canine diversity test are well aware of the efficacy of the test they offer – or they would not have developed it, published peer-reviewed studies using it, nor would VGL sell it.
So here are the important differences. Unlike the UC Davis test, the Embark test is designed to look at well known regions of the genome that are non-neutral or “adaptive,” areas “shown to harbor deleterious alleles” as Chu says. These are of interest to Embark because their stated mission is to further research looking for new disease traits, and that’s perfectly appropriate, since the Embark test is a diagnostic screening test. Their chip is the right tool for that job and their ample markers should help to find specific mutations.
These regions, however, are inherently subject to selection, because expressed traits in purebred dogs are selected for and against by humans. This is what causes runs of homozygosity. Genes for color and coat in a single breed can be the same in all dogs due to human selection. Disease genes, by contrast, are actively selected against by breeders, so ideally dogs are homozygous for healthy genes. These regions therefore can only offer a skewed picture of multi-locus heterozygosity or estimates of inbreeding. That makes them less appropriate for assessing genetic diversity, genetic drift and population structure.
To illustrate more clearly, imagine if we were to look at the genes that control coat type only. In such a case, Poodles of all varieties, Irish Water Spaniels, Portuguese Water Dogs and Komondors would all look highly related because they all have the curly coat genes. These breeds however are all genetically distinct. The breeds would also look highly inbred, since curly coats require a homozygous pair of alleles – but these dogs and breeds may or may not be inbred at all. This is why the specific regions sampled in any panel matter.
In order to best assess population structure, conservation and population geneticists have long used what are called “putatively neutral” microsatellites – or STRs. These markers are selected precisely because they cannot be selected for or against, and they are well known for being very informative for showing subtle differences in relatedness. What’s more, STRs are better at assessing recent genetic relatedness than SNPs due to their more rapid mutation rates, while SNPs are better at assessing ancient relatedness because of their very slow mutation rate. Because the changes in dog breeds that are of the most importance to breeders have happened in the last 150 years, STRs are an appropriate tool for our task as dog breeders. STRs are so powerful in fact that with only 13 STRs, it’s possible to positively identify any human on earth, and with that same number confirm paternity or other relationships.
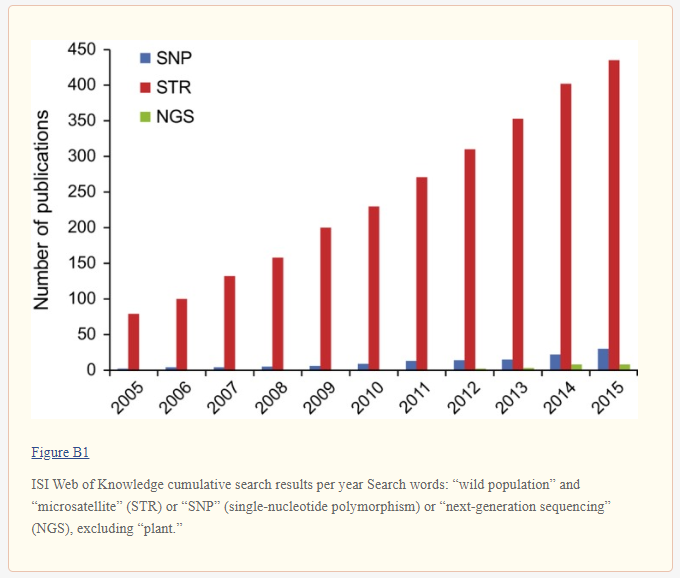
Meanwhile, Embark would have breeders believe that microsatellites are somehow no longer in use, knowing that breeders will not bother or know how to find out if that’s true. It’s not at all true. Here’s a large study on white-tailed deer from just a few months ago that describes 58 projects studying these deer and using microsatellites. Here’s another recent paper on elephant seals that discusses genetic bottlenecks detected by microsatellites. Here’s another recent paper discussing the mapping of microsatellites for the first time in camels for eventual use in genetic diversity studies. And here’s a study on Galapagos giant tortoise species that shows that poorly designed and interpreted SNP panels, though numerous in markers, can be less than accurate in showing population structure (how families in a population are related) if not done well. It also showed how a much larger subsequent SNP panel, properly designed, confirmed the accuracy of previous studies on population structure in these tortoises that used only 10 microsatellites.
This same methodology also accurately shows genetic relatedness between dogs and dog breeds, not just by VGL but by others as well, as shown in this paper published this year. Purebred dogs are more inbred than humans, so while some microsatellite panels validated this year require only 13 carefully selected STR to determine canine breed status, the VGL uses a larger panel which is adept at managing even very inbred breeds like Doberman Pinschers. What’s more, their 33 markers have been proven by VGL to offer the same accuracy in assessing relatedness as 58 markers do. When using STRs in conservation algorithms, the power is not in the number of markers, it’s in the number of alleles, or versions, at each marker. Relatedness calculations are based on probability, and so there’s a diminishing rate of improvement in accuracy as any new marker is added – kind of like the odds of getting heads when flipping 100 coins is the same as when flipping 1000 coins. (You can read more about relatedness estimators here.) Once you get to a high probability conclusion, additional markers simply do not materially improve that conclusion. That’s why human paternity tests never definitively say someone is the father of a child – they always say there’s a 99.999% chance someone is the father. Adding markers to the test would only make the probability 99.99999% likely to be the father.
Here’s another thing Chu doesn’t mention – there is no legitimate way to compare what we at BetterBred do with the VGL canine diversity test and what Embark does with their runs of homozygosity on their Illumina chip. They are vastly different analyses, and there is no scientific reason the number of STR markers in the VGL test used in our assessment should be compared one to one to the number of SNP markers in Embark’s test. The use of IR (the internal relatedness calculation) based on microsatellite data has been peer reviewed and published innumerable times in 18 years of scientific literature – and if Embark prefers to use another method, that’s fine, but there is no excuse for this constant harangue – only a profit-driven intent.
Ultimately the numbers game in SNPs is purely a marketing message that is put out by Embark to capture market share among breeders. This fact is best illustrated by comparing their 230,000 SNP test not to the VGL test, but rather to Mars Petcare’s Optimal Selection, also called MyDogDNA in Europe. Mars uses SNPs as well, and offers breeders the exact same number and accuracy of diagnostic, trait and inbreeding information as Embark, and has a better breeding tool, but the Mars test uses only 2,642 SNPs. If numbers mattered so much, that wouldn’t be possible.
Yet Embark never, ever goes after Mars or insults their comparatively few SNP markers, perhaps because Mars is a huge conglomerate with whom they really can’t compete. Instead, they choose to go after BetterBred, a tiny little company with a growing user and fan base, and the not-for-profit Veterinary Genetics Laboratory, a world leading academic and research institute that (conveniently for Embark) simply does not engage in commercial competitiveness. They back up their standing with research, not marketing blogs or private sales-oriented consultations like the ones Chu offers.
As for the content of Chu’s statement, it’s almost too easy to refute. Chu and her colleagues at Embark, none of whom seems to be a breeder or specializes in breeding, apparently do not understand that there is more to breed management than estimating inbreeding. They also seem not to understand that breeding solely for heterozygosity – no matter how it is estimated and in any breed – is foolish. Aaron Sams’ blog post referenced by Chu and comparing the VGL and Embark tests for breeding is poorly done, misleading, and does not inspire confidence in the long awaited Embark breeding tool. It is surprising that no one at Embark caught the many holes in it. That Chu would highlight that post means she didn’t catch them either. We certainly did. Here’s a bit of what we caught.
First, Sams does not describe the population of dogs he uses in his simulation – not the breed, not the number of dogs, not the allele frequencies or alleles per locus, not the original relatedness of the individuals, all of which can significantly affect the outcomes of any breeding program. If this were an area of expertise for Sams, he would know that different populations will have different outcomes, and purebred dog breeds vary widely in genetic diversity and relatedness and are never generic. For his simulation to be taken seriously, these pieces of information should be included.
Second, Sams sets his criteria very strangely:
- His first criterion is pedigree relatedness based on two generations, which is a bit shocking. Experienced breeders and researchers know two generation pedigrees are meaningless in purebred dogs for assessing relatedness or inbreeding in breeding programs.
- His second criterion is haplotype heterozygosity using not 230,000 SNPs, but in fact 50 separate haplotypes derived from Embark’s test. This is normally how genetic software packages do this of course – and 50 loci is really not much more than 33 loci – which sort of ruins Embark’s “more is more” argument.
- His third is genetic relatedness calculated using those 50 haplotypes – though we don’t know how it is calculated or why the calculation used was chosen.
- His fourth criterion is heterozygosity in “33 loci positioned across the genome in a manner similar to UC Davis’ STR panel” – meaning not the same carefully selected and validated STR loci used by VGL – just similarly positioned fictional ones.
- His fifth is the internal relatedness calculation using his simulated STRs, which is, in essence, another estimation of inbreeding and not comparable to genetic relatedness calculations.
Third, Sams cannot compare his company’s product with anything done by BetterBred because he hasn’t used our method and clearly knows nothing about it. Nobody, not BetterBred nor VGL, recommends selecting mates based on heterozygosity in polymorphic STRs for reasons that should be obvious to anyone with some experience, or regular readers of our blog. Moreover, simply selecting for lowest average IR in offspring is not something we at BetterBred recommend when selecting mates, also for reasons that should be obvious to an expert, and is well known to our users. We opted for more effective ways to preserve allelic richness in a population, which ultimately results in higher average genome wide heterozygosity in dog breeds.
Fourth, Sams’ ultimate conclusion is that selecting mates using genome-wide heterozygosity and genome-wide relatedness derived from Embark’s SNPs will conserve 0.5% more genetic diversity (allelic richness) over 20 generations than by using 2 generation pedigrees, 4.3% more than STR heterozygosity and 1% more than using internal relatedness calculated with STR data. He also concludes that using two generation pedigrees alone will conserve 0.8% more genetic diversity than STR internal relatedness. Sams claims that “in all comparisons of an STR model to any other model” the statistical significance had an amazing P value of < 10-48 , even though the mean value for the random mate selection and STR internal relatedness selection are virtually identical and their deviations are overlapping. (There’s definitely something fishy about that math.) Furthermore, while the graphics are specifically designed to make these results look dramatic, these are only very negligibly different outcomes (look at the actual numbers on the vertical axis) – especially considering the 20 generation time frame.
In essence, all of the methods in this simulation involve loss of about 25% of genetic diversity after 20 generations. In Sams’ zeal to prove his tool superior to VGL’s, he botched the VGL comparisons, and has really only legitimately shown that his company’s breeding tool barely competes with the use of two generation pedigrees! Given that there are dozens of peer reviewed publications that show genetic data is more accurate than pedigree data, all this blog post really does is sow doubt in Sams’ dataset or his analysis.
There is more that is easy to critique in Sams’ post, but what remains most astounding, is that he proves, on his own, unsolicited, and Embark happily publishes and promotes (!) that the Embark breeding tool is essentially of no more use to breeders than two generation pedigrees, which breeders already have. Hardly a ringing endorsement.
It is ironic that Sams ends this very flawed post saying they are working hard on finding a test that will really help breeders, who should rely on “expert assistance from population geneticists and veterinarians.” Happily, the eminent population geneticists and veterinarians at UC Davis VGL who designed the STR based canine diversity test have already offered breeders such a test, have validated it in nearly 30 breeds, have published 5 peer reviewed papers using these panels, and are working on more breed analyses constantly. Perhaps it is their consistent, methodical success that makes Embark determined to undermine breeders’ confidence in their careful scholarship and the extensive help they offer breeders.



